hicInterIntraTAD¶
Extracts and computes different inter and intra TAD values and ratios.
usage: hicInterIntraTAD [--matrix MATRIX] [--tadDomains TADDOMAINS]
[--outFileName OUTFILENAME]
[--outFileNameRatioPlot OUTFILENAMERATIOPLOT]
[--fontsize FONTSIZE] [--dpi DPI] [--threads THREADS]
[--help] [--version]
Required arguments¶
- --matrix, -m
The matrix which was used to compute the TADs
- --tadDomains, -td
The TADs domain file computed by hicFindTADs.
- --outFileName, -o
Outfile name
Default: “output_interintra_tad.tzt”
Optional arguments¶
- --outFileNameRatioPlot, -op
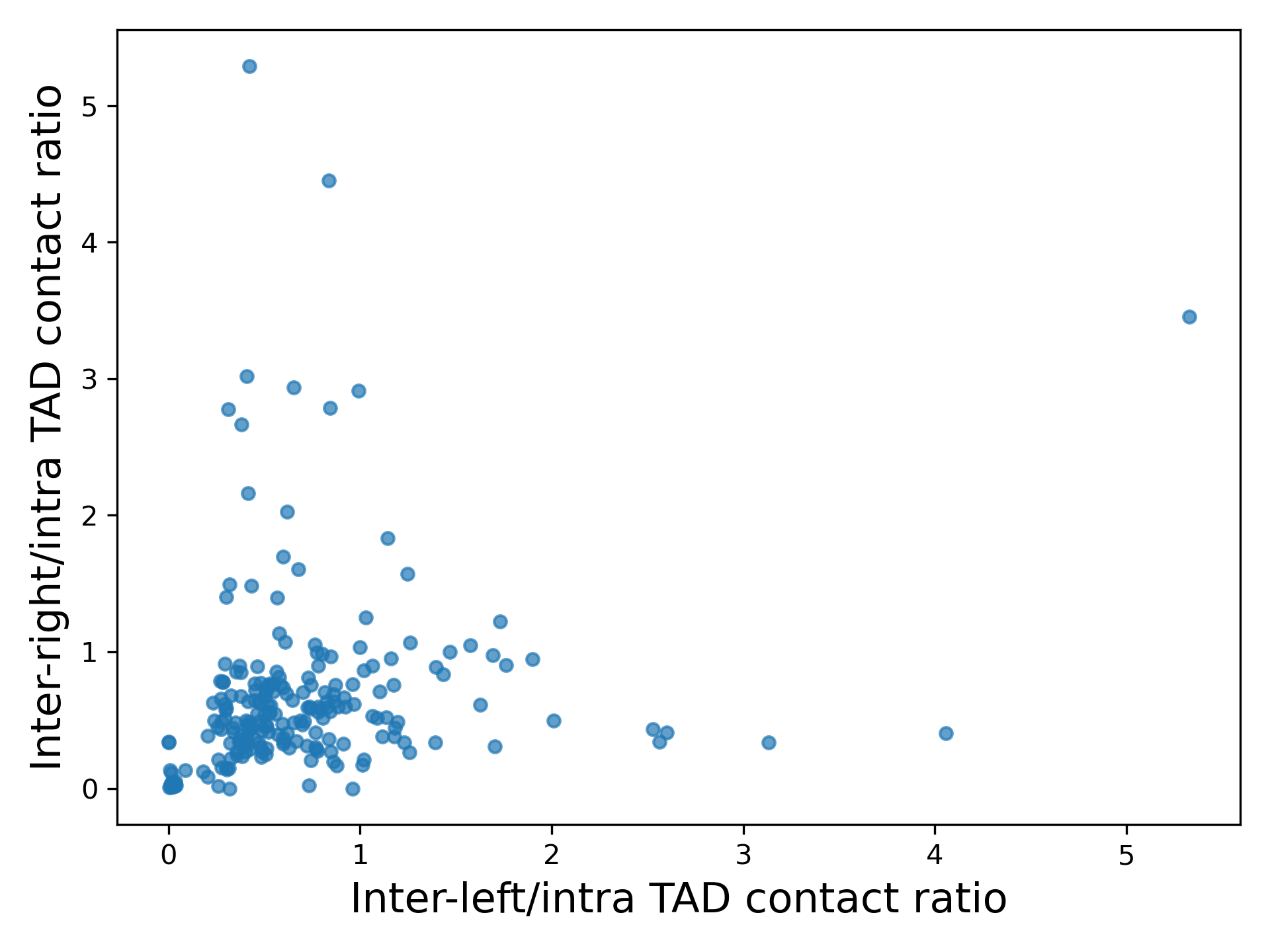
Outfile name for the inter-left/intra vs inter-right/intra ratio plot
Default: “ratio.png”
- --fontsize
Fontsize in the plot for x and y axis.
Default: 15
- --dpi
The dpi of the scatter plot.
Default: 300
- --threads, -t
Number of threads to use, the parallelization is implemented per chromosome (Default: 4).
Default: 4
- --version
show program’s version number and exit
This tool computes and extracts for given TADs the inter-left, inter-right and intra-TAD data: the absolute number of contacts, the density (non-zero contacts / all possible contacts) and the ratio of inter-left/intra, inter-right/intra and (inter-left + inter-right)/intra TAD contacts. The data is saved in one file where it can be used for further, user-specific computations outside of HiCExplorer. Also a scatter plot of the ratio inter-left/intra vs. inter-right/intra is created.
# Created with HiCExplorer's hicInterIntraTAD version 3.7-dev
# Chromosome start end name score strand inter_left_sum inter_right_sum inter_left_density inter_right_density inter_left_number_of_contacts inter_right_number_of_contacts inter_left_number_of_contacts_nnz inter_right_number_of_contacts_nnz intra_sum intra_number_of_contacts intra_number_of_contacts_nnz intra_density inter_left_intra_ratio inter_right_intra_ratio inter_left_inter_right_intra_ratio
chr1 4400000 6200000 ID_0.01_1 -0.5630275 . 0 265.6981737879304 0 1.0 0 180 0 180 780.0186987409819 324 324 1.0 0.0 0.340630518494993 0.340630518494993
chr1 6200000 7300000 ID_0.01_2 -0.235798 . 288.00513572726237 327.503479611623 1.0 1.0 198 231 198 231 339.91508704783513 121 121 1.0 0.8472855330682405 0.9634861531331044 1.8107716862013452
chr1 7300000 9500000 ID_0.01_3 -0.44334 . 340.12385944568155 159.40880484157745 1.0 1.0 242 110 242 110 1078.1133262629996 484 484 1.0 0.31548061892958207 0.14785904316211984 0.46333966209170185
chr1 9500000 10100000 ID_0.01_4 -1.021538 . 186.65816676710497 124.2235190772146 1.0 1.0 132 84 132 84 118.58286349492002 36 36 1.0 1.5740737005823884 1.0475672067282826 2.6216409073106712